www.accnweb.comwww.acolytebiomedica.comwww.affiniti-res.comwww.allscience.cowww.arachnoiditis.infowww.aralbio.comwww.arexdb.orgwww.aureus-pharma.comwww.autotestvih.infowww.aviabiosystems.comwww.axis-shield-density-gradient-media.comwww.axonscientific.comwww.bacterialgenomics.orgwww.biochempages.comwww.biomeeter.comwww.bioprodmag.comwww.biosantech.orgwww.blog-biosyn.comwww.camb-labs.comwww.camelgate.comwww.ceterix.comwww.clcngs.comwww.cornellcelldevbiology.orgwww.crocgenomes.orgwww.designmedix.comwww.diagxotics.comwww.dnachip.orgwww.dorbiopharma.comwww.dpgp.orgwww.eaa2020.orgwww.elsf.orgwww.elsf.orgwww.eurechinoreg.orgwww.eurowestnile.orgwww.ewctrainingcenter.orgwww.extinct-website.comwww.fluidsforlife.comwww.gcmsservice.comwww.genemol.orgwww.gensetoligos.comwww.immunopred.orgwww.inserm-actualites.comwww.interchromforum.comwww.jhindia.comwww.kansasbio.orgwww.lakesyn.comwww.leishnet.netwww.ligand.infowww.luxan.co.ukwww.maizemap.orgwww.nabfa-blackfly.orgwww.nakedbiome.comwww.ncnsd.orgwww.neurostemcell.orgwww.neusilin.comwww.niddkrepository.orgwww.noabbiodiscoveries.comwww.novactabio.comwww.novexin.comwww.ohmxbio.comwww.omicsbio.orgwww.ondex.orgwww.orengogroup.infowww.perceptivebioscience.comwww.pharma-planta.netwww.phenyx-ms.comwww.plantnames.orgwww.preclinomics.comwww.procellbiotech.comwww.proteincrystallography.orgwww.proteinforest.comwww.qcmg.orgwww.qhek.comwww.reiclabs.comwww.reseqtb.orgwww.sebio.orgwww.smb2019.orgwww.surechem.orgwww.tankfishtips.comwww.theebi.orgwww.treatgene.comwww.trpchannel.orgwww.visiblegenetics.comwww.wheaton-uk.comwww.whybiotech.comwww.wolfelabs.comwww.younginfrontier.com
Scientists Identify Corona virus Strain
Scientists Identify Corona virus Strain In Cyprus-‘Deltacron’-That Blends Delta And Omicron
Scientists in Cyprus have found 25 cases of a strain of the coronavirus that they say combines elements of the delta and omicron variants, dubbing it “deltacron,” with a high proportion of the variant found in patients hospitalized for Covid-19, a professor involved in the identification of the new strain said Saturday.
People at the Larnaca International Airport in Cyprus (Photo by CHRISTINA ASSI/AFP via Getty Images)
KEY FACTS
- The discovery was made by Leondios Kostrikis, professor of biological sciences at the University of Cyprus and head of the Laboratory of Biotechnology and Molecular Virology, and his team, Bloomberg reported.
- The new strain has “omicron-like genetic signatures within the delta genomes,” according to Bloomberg.
- Deltacron cases were found in 25 samples taken in Cyprus, of which 11 were from patients hospitalized with Covid and 14 from the general public, according to Cyprus Mail, a local English daily.
Analysis shows deltacron is more often found in patients hospitalized with Covid-19 than those with Covid-19 who are not hospitalized, Kostrikis said.
It is “quite possible” that the new strain has not been found elsewhere, and the sequences of the cases have been sent to GISAID, a Germany-based international database that tracks developments in the coronavirus, the Cyprus Mail reported.
Australia’s Last Zero‑CovidHoldout Cancels Border Reopening Plans As Omicron Surges Across Country
KEY BACKGROUND
In December 2021, Paul Burton, Moderna’s chief medical officer, told the U.K.’s House of Commons that the co-existence of delta and omicron increased the chances of a new variant as a result of them trading genes, the Daily Mail reported. Such recombination is common in coronaviruses, according to the New York Times. Several studies have suggested recombination could cause the virus to change in “dangerous ways,” but could help researchers develop drugs to treat the virus.
Some experts suggest omicron variant
Some experts suggest omicron variant may have evolved in an animal host
When COVID-19 variants arise, the accepted wisdom is that the constellation of mutations they contain developed in an immunocompromised person who contracted the virus and couldn’t shake the infection. But some scientists have an alternative theory for where the latest variant of concern, omicron, may have acquired the unusual mutations that stud its spike protein.
They speculate the virus could have evolved in another animal species.
The theory goes that some type of animal, potentially rodents, was infected with the SARS-CoV-2 virus sometime in mid-2020. In this new species, the virus evolved, accumulating roughly 50 mutations on the spike protein before spilling back over into people.
Kristian Andersen, an immunologist at the Scripps Research Institute, is among those who has been raising the idea that Omicron may have emerged from a reverse zoonotic event.
(A zoonotic event is when an animal pathogen starts to infect and spread among people. A reverse zoonosis is when such a virus passes back into an animal species.)
“I know that most people think that these [come from] immunocompromised individuals, and I do think that that’s plausible, but to be perfectly honest, I actually think this reverse zoonosis followed by new zoonosis seems more likely to me given just the available evidence of the really deep branch, and then the mutations themselves, because some of them are quite unusual,” Andersen told STAT.
- “I don’t think we should dismiss that possibility, because I think it’s definitely on the table.”
- A number of other scientists who study the evolution of viruses have told STAT they think the idea isn’t out of the question. Some place more weight on the theory that variants develop in immunocompromised people, while others feel there isn’t enough evidence at this point to favor one option over the other.
- “Personally, I think it’s probably more likely it was circulating undetected, in an immunocompromised individual,” Emma Hodcroft, a molecular epidemiologist at the Institute of Social and Preventive Medicine in Bern, Switzerland, said via email. Having said that, though, Hodcroft insisted that it is important to explore the hypothesis.
- “I would certainly consider it a plausible alternative hypothesis to the evolution during a persistent infection in a human,” said Andrew Rambaut, a professor of molecular evolution at the Institute of Evolutionary Biology in Edinburgh. He cautioned that coming up with a definitive answer won’t be quick.
- “I am not sure we will be in a position to say for sure for a while,” Rambaut wrote in an email.
One of the peculiar traits of SARS-2 underpins this thinking. It is what virologists describe as a promiscuous virus; it is capable of infecting a number of species. Dogs and house cats. Large cats. Mink. White-tailed deer. Given how easily the virus seems to jump from species to species, people studying it assume this list will grow.
The original virus that came out of Wuhan, China, in early 2020 did not infect rodents. But as variants — Alpha, Beta, Delta — started to emerge, those viruses could infect rodents.
Robert Garry, a professor of microbiology and immunology at Tulane Medical School, has been tracking the SARS-2 mutations that have arisen. Seven are associated with rodent adaptation — the changes that seemed to allow the virus to infect mice, rats, and related species. All seven of those mutations are in omicron, Garry noted. He believes it’s a toss-up whether the variant developed in an animal or a human host, but if it’s the former, his bet would be on rodents.
Getting a firm answer might require enormous luck. Scientists are looking at various animal species to see if they can be infected with SARS-2; were they to find viruses like omicron in any, that would swing the needle.
But Michael Worobey, a professor of evolutionary biology at the University of Arizona, thinks one could do some experiments on selected species of wild animals to see if they can be infected and if, when infected, similar patterns of viral evolution occur.
Studying the molecular clock of viruses that spread in animals — looking at the speed at which they evolve and comparing it to SARS-2 evolution in humans — could also provide some clues, said Worobey, who initially thought Andersen’s idea was not impossible, but not the likeliest of explanations for omicron. After hearing details of the explosive outbreak in white-tailed deer, he’s rethinking the idea.
For Worobey, the question is whether any animal species can become chronically infected with SARS-2 — in effect, whether there are animal species in which SARS-2 lingers in the way it does in immunocompromised people. That could put positive selective pressure on the virus — in other words give it an incentive to mutate to stay ahead of the animal’s immune response.
“It does move my thinking in terms of omicron possibly having come from a reservoir, if there are [animal] reservoirs that do chronic infections,” he said.
Part of what leads Andersen to wonder about an animal source is the fact that the variant traces back to viruses that were spreading over a year ago. “That in itself you need to be able to explain,” he said.
Angela Rasmussen, a coronavirus virologist at the University of Saskatchewan’s Vaccine and Infectious Disease Organization, agreed.
“I think it’s pretty obvious to everybody … that this virus has been on an independent evolutionary track for quite some time and it’s very surprising, which to me just kind of goes back to say well, the idea that this could be … plausible,” she said.
Regardless of whether this variant emerged in another species or not, given SARS-2’s ability to jump species, it is possible the world will face animal-derived variants in the future, Garry warned. The upshot of that? “We’re going to have to keep tweaking the vaccines.”
Bio-informatics ressources
| [GdmCl] and [Urea] from refractive index by Sosnick lab web |
| 1. Compute pI/Mw |
| 16S rRNA database and workbench (most recently updated data can be downloaded here) |
| 2. ProtParam |
| 3. Translate |
| A beta word processor run over the web |
| A command-line tool that generates a table of several useful geometric measurements for each residue or base from a PDB file. |
| A database of research protocols in a variety of life science fields. It has a popular discussion forum. |
| A DNA part, plasmid, microbial strain, and Arabidopsis Seed online repository with physical sample tracking capabilities |
| A free all-in-one platform for intuitive vector design and automatic assembly. Supports main construction techniques, manual and auto sequence annotation, alignment, primer design and compatible to all file formats. |
| A free DNA sequence viewer and annotation tool (Java based). The Sanger Center also develops a number of other miscellaneous tools for download |
| A helper application for your web browser that allows you to view 3-dimensional structures from NCBI’s Entrez retrieval service. It doesn’t read PDB files but can be more straightforward to use than DeepView. |
| A number of web tools and calculators to assist in the design and execution of molecular biology research |
| A powerful, free online & downloadable genetic engineering all-in-one platform for molecular & synthetic biologists |
| A site where you can explore the various features of the JBEI Registry software, and even get some work done! |
| A tool which allows the computation of the theoretical pI (isoelectric point) and Mw (molecular weight) for a list of Swiss-Prot and/or TrEMBL entries or for user entered sequences. |
| A tool which allows the computation of various physical and chemical parameters for a given protein stored in Swiss-Prot or TrEMBL or for a user entered sequence. |
| Access to sequence-validated, full-length, protein-coding, mammalian cDNA clones |
| AddGene Vectors can be directly imported into Genome Compiler software platform, in which you can easily edit and visualize it. |
| Addgene is a a non-profit plasmid repository where scientists can archive and share their plasmids. Addgene assists with data submission and all tech transfer issues. Plasmids can be requested from Addgene for a fee to cover expenses. |
| Addgene’s Vector DB |
| Affymetrix Tools software |
| AGADIR by Serrano lab web |
| All-in-one molecular cloning and genetic engineering design, simulation & management tool for complex synthetic biology and metabolic engineering project. |
| An algorithm to predict the helical content of peptides. |
| An incomplete web version of the publication Escherichia coli and Salmonella: Cellular and Molecular Biology. (Subscription required) |
| An integrated tool for PCR primers or probe design, in silico PCR, oligonucleotide assembly and analyses, alignment and repeat searching |
| An integrative framework for discovering functional relationships among proteins. |
| An open-source java program for modeling gene regulatory networks. Users can rapidly build networks by specifying their topology, initial conditions, connectivity, and known parameters. Ingeneue can then search/explore paramter space for desired behavior, simulate the effects of noise and mutation, and generate statistics/time graphs of the system. |
| Antibody Resource |
| ApE A plasmid Editor software |
| Appendix by Ambion, Inc. |
| Apps for predicting RNA and DNA folds, calculating Tm’s and free energies. Runs mfold + UNAfold servers |
| Artemis by the Sanger Center software |
| AVID by Keating lab web |
| Awesome program for viewing and studying protein structure. |
| Backbone-dependent rotamer library by Dunbrack lab web |
| Barrick Lab of UT Austin |
| BenchFly |
| Benchling web |
| Beta suite of office applicatins offered over the web |
| BioCyc |
| BioEdit software |
| Bioinformatics Toolbox from DNA2.0 web |
| BioNumbers |
| BLAST web |
| both prokaryote and eukaryotic promoter prediction |
| BRENDA |
| Calculate a number of parameters for your nucleotide polymers |
| Calculator to determine a protein’s contact order |
| Calculators |
| CambridgeSoft software |
| Cancer Genome Anatomy Project web |
| CDC Disease Conditions |
| CLC Sequence Viewer software |
| Cn3D by NCBI web |
| Colibri by Institut Pasteur |
| Collaborative encyclopedia of biodiversity |
| Collection of links to many pages to calculate parameters of your favorite proteins |
| Collection of online tools for codon optimization and shuffling, restriction site editing, and so on. |
| Collection of science-related links, including links to journals, catalogs, and tools. See their about page. |
| Collection of thousands of pathway/genome databases for many organisms, plus software tools for understanding their data |
| Combine genetic building blocks by drag-and-drop, codon optimize, restriction site editing, sequence oligo design etc. See BMC Bioinformatics 2006 Jun 6;7(1):285 for more detail. |
| Community for sharing and discussion of research papers |
| Comprehensive biochemical pathway and gene function site for E. coli |
| Comprehensive enzyme information system |
| Computational server for finding/calculating summary data about a wide range of topics, plus useful widgets like reagent tables and gene lookup |
| Consensus structural annotations and 3D models for sequences of model organisms (built upon numerous other useful related resources) |
| ConSurf |
| Contact Order Calulator by Baker lab web |
| Contains many Javascript tools to do common tasks. |
| Control translation initiation rate and predict protein production levels |
| Control translation initiation rate and predict protein production levels. |
| Coredemia web |
| Crowdfunded scientific research |
| Current Protocols in Molecular Biology |
| CyberCell Database (CCDB) by Institute for Biomolecular Design |
| Dang by Richardsons’ lab software |
| Database of all E. coli genes and sequences (Entrez data is pulled from here) |
| DeepView by GlaxoSmithKline & Swiss Institute of Bioinformatics software |
| Design genes de novo with a powerful and intuitive user interface |
| Design Zinc Finger DNA binding proteins |
| DeviceEditor: a visual DNA design canvas that serves as front-end for j5 |
| Digital lab notebook for organizing and sharing experiments, findings, protocols, and more. |
| DNA |
| E. coli genome browser; get sequences, see the position of your gene in the chromosome, see the function of your gene, and other fun stuff. You can also search for protein sequences/motifs within the E. coli genome. |
| Easy-to-use Bioinformatics manipulation tools for UCSC data |
| EcoCyc |
| EcoGene |
| EcoSal by ASM Press |
| EMBL-EBI |
| ENCODE |
| Ensembl |
| Ensembl Genome |
| Enter a protein to search for antibodies and ELISA kits |
| Entrez (NCBI) |
| Eukaryotic genome browser |
| Evernote software app |
| ExPASy Proteomics server by the Swiss Institute of Bioinformatics web |
| Experiment web |
| Extensive database of protein sequence and functional information |
| FastPCR software |
| Filtered index of diseases from the CDC database |
| Finds rare codons in a coding sequence. |
| Finds regions of similarity between biological sequences. |
| Focused on protein crystallization but contains a lot of generally useful information about various reagents with respect to proteins. |
| Follow a link to the underlying open-source software source code. |
| Formerly curated by MIT, this is an open repository of BioBricks; the place for all your standard biological parts. |
| Free digital version of the paper Metabolic Pathways map. Because it is a PDF, it is also searchable. |
| Free to download and works on Mac or PC. User agreement is somewhat restrictive, i.e. you cannot sell genes designed using the tool without permission. |
| Free version of SnapGene. Allows mapping of DNA up to 1 Gb in length. Convenient viewing and annotation tools for genetic constructs. |
| Free, feature-rich drawing program with support for SVG and PDF images |
| Free/simple online toolset for sequence manipulation |
| Galaxy web |
| GdmCl and urea concentration calculator from index of refraction. |
| Gene Design by Boeke lab web |
| Gene expression in E. coli by Ehrmann lab |
| GeneDesigner by DNA 2.0 software |
| GeneDesigner by DNA2.0 software |
| General |
| General Search/Reference |
| Generates and annotates a plasmid map based on sequence data |
| Generates research article recommendations unique to each user. Site also has a special academic community. |
| GeneWarrior web |
| Genome and Metabolism |
| Genome Compiler web software |
| Genome Compiler web software |
| Genome Compiler web software |
| Genome3D |
| Genomic database geared towards high-level functions of the biological system |
| GETAREA by Sealy Center for Structural Biology web |
| Google Documents by Google web |
| GreenGenes |
| Growing collection of laboratory protocols and techniques |
| Handbook of protocols. (Links to Wiley Online Library) |
| Handbook of protocols. Subscription only. External link |
| High throughput sequence analysis (orders of magnitude faster than BLAST) and other software tools |
| High-level design, analysis and transmission of protein constructs. Pinecone matches users’ designs with CRO’s or DNA synthesizers to produce genetic starting material (dsDNA, plasmid or purified protein). Facilitates mutation and combinatorial protein sets that benefit from manufacturing economies of scale. |
| Huge, open access life science protocol repository for discovery and sharing of scientific methods |
| Hugely extensive collection of resources, databases, and tools related to diverse aspects of bioinformatics and molecular biology, often containing everything one might need |
| IDT SciTools web |
| iGEM Parts Registry web |
| Ingeneue by George von Dassow, Eli Meir, Edwin Munro, and Garret Odell at the Center for Cell Dynamics software |
| Inkscape web |
| InstaGrok software |
| Interactive simulations for science and math |
| Inventory of Composable Elements (ICE) – The public instance of the JBEI Registry |
| j5, DeviceEditor, and VectorEditor web |
| j5: DNA assembly design automation for (combinatorial) flanking homology (e.g., SLIC/Gibson/CPEC/SLiCE/yeast) and type IIs-mediated (e.g., Golden Gate/FX cloning) assembly methods |
| Jalview software |
| Java tools for creating RNA secondary structure diagrams |
| KEGG |
| Khan Academy (Biology) |
| Lab Techniques |
| Libraries of sidechain rotamer from protein structures |
| Light and intuitive tools for diverse molecular biology manipulations |
| Local cell colony counting software |
| MABL web |
| Mammalian Gene Collection |
| Massive, searchable rRNA database (especially strong for microbes) |
| MCDS software |
| Mendeley software app |
| MetaBase |
| Metabolic Pathways Poster PDF by SigmaAldrich |
| MetaCyc |
| Metazome web |
| MicrobeWiki |
| Microbial physiology |
| Microchip and SNP analysis software. DevNet Tools and other compatible programs also listed for download and use |
| Modeller by Sali lab software |
| Molecular Cloning by Sambrook and Russell |
| Molecular graphics system with an embedded Python interpreter designed for real-time visualization and rapid generation of high-quality molecular graphics images and animations. The latest version does not run on OSX 10.3. (from Kathleen). |
| Molecular visualization program for displaying, animating, and analyzing large biomolecular systems using 3-D graphics and built-in scripting. Generates pretty high resolution pictures of protein structures. |
| Multi-organismal member of BioCyc collection; catalogs entire universe of metabolism |
| NAS report on responsible conduct in research. |
| NEB Cutter by New England Biolabs, Inc. web |
| Numerous proprietary applications for research and composition (ChemBioDraw is usually accessible via academic institution email) |
| OligoCalc Oligonucleotide Calculator web |
| On Being a Scientist by National Academy of Science |
| Online apps for cloning, molecular biology, and analysis |
| Open source PCR primer design. Written in Perl/Tk. |
| OpenCFU software |
| Organize, annotate, search, and cite references from a PDF library (loads of extra features) |
| Origin software |
| OWW Materials |
| OWW Protocols |
| PAIRCOIL2 by Keating and Berger labs web |
| PaR-PaR allows researchers to use liquid-handling robots effectively, enabling experiments that would not have been considered previously. After minimal training, a biologist can independently write complicated protocols for a robot within an hour. |
| PaR-PaR Laboratory Automation Platform web |
| Parts list for functional elements in the human genome |
| PerlPrimer software |
| Permits writing of shareable, web-based text documents |
| PhET |
| Pinecone by Serotiny web |
| PlasMapper web |
| Polonies are colonies of PCR amplicons derived from a single molecule of nucleic acid. |
| Polony Protocols by Church and Mitra Lab |
| Powerful and straightforward local and cloud-based reference manager |
| Powerful but expensive vector editor. Supports most sequence types and easily manipulates genetic constructs. |
| Powerful cross-platform note-taking, composition, and organization with cloud syncing. |
| Powerful suite of open source bioinformatics tools for performing microbiome analysis from raw sequences. |
| Primer3 web |
| Program for homology or comparative modeling of protein three-dimensional structures by satisfaction of spatial restraints. |
| Program for multiple sequence alignment editing, visualization, and analysis |
| Promoter prediction web |
| Protein |
| Protein |
| ProteinProspector by UCSF Mass Spectrometry Facility web |
| Proteome-level phylogeny and genomics |
| Proteomics tools for mining sequence databases in conjunction with Mass Spectrometry experiments. |
| Protocol-online by Dr. Long-Cheng Li |
| Protocols.io app web |
| PubChase software app |
| PyMOL software |
| QIIME (Quantitative Insights Into Microbial Ecology) software |
| Qiqqa software |
| Quality-controlled, aligned and annotated Bacterial and Archaeal 16S rRNA sequences, and Fungal 28S rRNA sequences, and a suite of analysis tools |
| Quickly access a host of tools for analyzing and manipulating DNA, RNA, and protein sequences |
| Rare Codon Calculator (RaCC) by NIH MBI Laboratory for Structural Genomics and Proteomics web |
| RBS Calculator in Genome Compiler |
| RBS Calculator in Genome Compiler |
| ReadCube software |
| Reconstruct and analyze phylogenetic relationships between molecular sequences |
| Reference manager, similar to Mendeley or Evernote. Can enhance PDFs and search PubMed from within the app. |
| Replacement for mFold for predicting nucleic acid folding. Downloadable and some applications are available online also. |
| Research Tools |
| Resources and tools related to the characterization of cancer gene expression profiles |
| Ribosomal Database Project |
| RNA |
| RNA |
| RNA secondary structure prediction and design |
| Saccharomyces genome database (yeast genome) |
| Science Gateway web |
| ScienceHack |
| Scientific graphing and data analysis software |
| Search all NCBI databases |
| Search engine for science videos with a review system |
| Search of rare codons in nucleotide sequence by “Practical Molecular Biology” web |
| Search, view, upload, create, and host scientific protocol videos |
| Selection of science videos and short courses, including the popular Crash Course series |
| Sequence file format converter from NIH web |
| Sequence Manipulation Suite web |
| Sequence reference for a large number of genomes |
| Serial Cloner software |
| Server for the identification of functional regions in proteins |
| Several tables describing statistical data on E. coli compiled from several sources. |
| Sfold web |
| SGD |
| Sharable, web-based word processing, spreadsheets, and presentations |
| Sharable, web-based word processing, spreadsheets, and presentations (alternative to Google Docs) |
| SIDD web |
| SILVA |
| Simple tools for plasmid design and manipulation |
| Sister to Ensembl, not limited to Eukaryotes |
| SnapGene Viewer software |
| Software environment enabling users to make a large number of bioinformatics analyses, combined with smooth data management, graphical viewing, and output options. |
| Solvent accessible surface areas, atomic solvation energies, and their gradients for macromolecules |
| Statistical Folding and Rational Design of Nucleic Acids. Predicts accessible RNA sites |
| Statistics to use from Saint John’s University. web |
| Stress induced DNA duplex destabilization. Finds destabilized sites in superhelical DNA. |
| Student edited resource on microbes and microbiology (curated pages reviewed by microbiologists) |
| Teaching |
| Technical suppport on protein crystallization by Hampton Research |
| The Complete set of tools for sequence viewing, annotation and alignment. Free online & downloadable and supports all file formats |
| The database of biological databases – External link |
| The database of useful biological numbers. (link to database) |
| ThinkFree Office Online by ThinkFree web |
| Tips and information on gene expression in E. coli |
| Tons of helpful resources like protocols, guides, links, and tools for lab work |
| Tool for finding restriction sites, et cetera. |
| Tool for seeking, collecting, and mapping information |
| Tool that lets you pick & evaluate primers from a DNA sequence |
| Tool to predict the parallel coiled coil fold from sequence using pairwise residue probabilities. |
| Tools and applications to aid in research tasks. |
| Translate is a tool which allows the translation of a nucleotide (DNA/RNA) sequence to a protein sequence. |
| Tree of Life |
| Try out the integrated online tools, including DNA sequence editing and annotation (Vector Editor) and auto-aligning sequencing trace files against a template. |
| UCSC Genome Bioinformatics |
| UNAfold by Michael Zuker. web |
| UNAFold web |
| UniProt |
| USEARCH software |
| Useful information for making or obtaining reagents, enzymes, buffers, etc. |
| VADLO is a search engine for Life Sciences Protocols, Online Tools, Databases, Software, and Biomedical Powerpoint Lectures. It also has Daily research cartoons, called *”Life in Research” Cartoons. |
| VADLO Search Engine |
| Vector NTI software |
| VectorEditor: a visual DNA editing and annotation tool |
| Vectors |
| Vectors |
| Vessel web app |
| Vienna RNA software |
| VMD by Theoretical and Computational Biophysics Group at UIUC software |
| Web tool for converting between sequence file formats. |
| Website with many useful nucleic acid parameters. |
| Windows-only sequence alignment editor (no longer maintained, but free to download) |
| Wolfram Alpha |
| WriteBoard by 37signals web |
| Writely by Upstartle, LLC. web |
| Writing/Composition/Organization |
| XRNA software |
| Zinc Finger Tools by Barbas lab web |
| Zoho web |
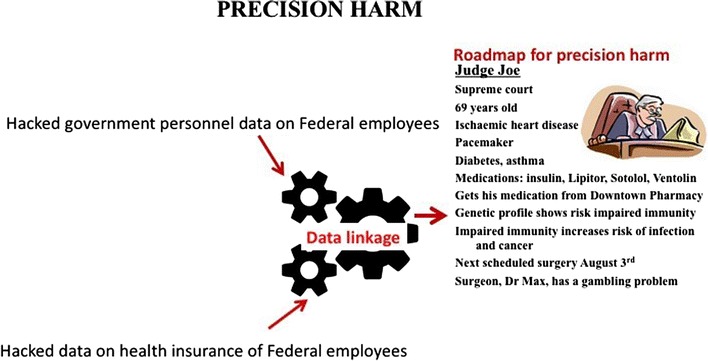
Converging and emerging threats to health security.
Advances in organic sciences have outpaced regulatory and authorized frameworks for biosecurity. Simultaneously, there was a convergence of scientific disciplines corresponding to artificial biology, information science, superior computing and many different applied sciences, which all have functions in health.
For instance, advances in cybercrime strategies have created ransomware assaults on hospitals, which may cripple health methods and threaten human life. New sorts of organic weapons which fall outdoors of conventional Cold War period considering may be created synthetically utilizing genetic code.
These convergent trajectories are dramatically increasing the repertoire of strategies which can be utilized for profit or hurt. We describe a brand new threat panorama for which there are few precedents, and the place regulation and mitigation are a problem. Rapidly evolving patterns of know-how convergence and professionalliferation of dual-use dangers expose insufficient societal preparedness.
We define examples within the areas of organic weapons, antimicrobial resistance, laboratory safety and cybersecurity in health care. New challenges in health safety corresponding to precision hurt in drugs can not be addressed throughout the remoted vertical silo of health, however require cross-disciplinary options from different fields.
Nor can they can’t be managed successfully by particular person nations. We define the case for brand new cross-disciplinary approaches in threat evaluation to an altered threat panorama.
A Bridge to the World.
Zhi-Hong Xu is a plant physiologist who studied botany at Peking University (1959-1965). He joined the Shanghai Institute of Plant Physiology (SIPP), Chinese Academy of Sciences (CAS), as a graduate pupil in 1965. He recollects what has occurred for the institute, through the Cultural Revolution, and he witnessed the spring of science finally coming to China. Xu was a visiting scholar on the John Innes Institute and within the Department of Botany at Nottingham University within the United Kingdom (1979-1981).
He turned deputy director of SIPP in 1983 and director in 1991; he additionally chaired the State Key Laboratory of Plant Molecular Genetics SIPP (1988-1996). He labored as a visiting scientist within the Institute of Molecular and Cell Biology, National University of Singapore, for 3 months every year (1989-1992).
He served as vp of CAS (1992-2002) and as president of Peking University (1999-2008). Over these durations he was closely concerned within the design and implementation of main scientific tasks in life sciences and agriculture in China. He is an academician of CAS and member of the Academy of Sciences for the Developing World. His scientific contributions primarily cowl plant tissue tradition, hormone mechanism in growth, in addition to plant developmental response to atmosphere.
Zhi-Hong, as a scientist and chief who has made an influence locally, known as up loads of glorious younger scientists returning to China. All of his efforts have promoted the quick growth of China’s plant and agricultural sciences. Expected remaining on-line publication date for the Annual Review of Plant Biology, Volume 71 is April 29, 2020. Please see http://www.annualreviews.org/page/journal/pubdates for revised estimates.
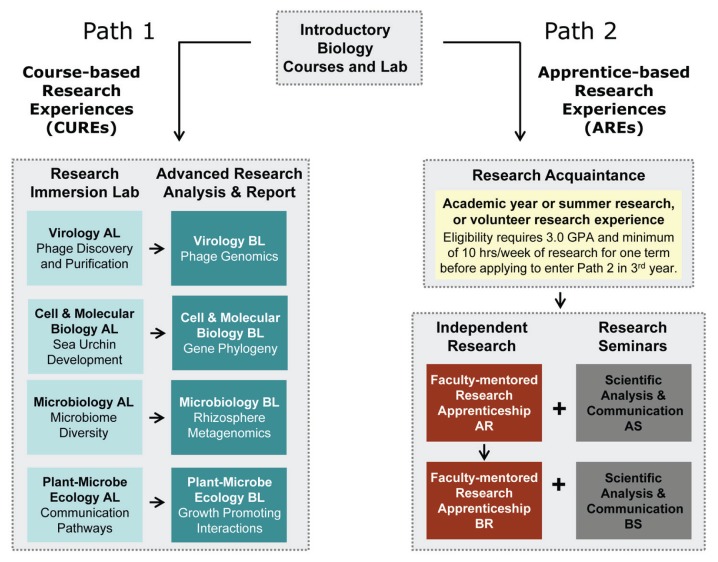
Comparing the Impact of Course-Based and Apprentice-Based Research Experiences in a Life Science Laboratory Curriculum.
This four-year research describes the evaluation of a bifurcated laboratory curriculum designed to offer upper-division undergraduate majors in two life science departments significant publicity to genuine analysis. The timing is crucial because it gives a pathway for each immediately admitted and switch college students to enter analysis.
To fulfill their diploma necessities, all majors full one of two paths in the laboratory program. One path immerses college students in scientific discovery skilled by means of workforce analysis initiatives (course-based undergraduate analysis experiences, or CUREs) and the different path by means of a mentored, impartial analysis venture (apprentice-based analysis experiences, or AREs). The bifurcated laboratory curriculum was structured utilizing backwards design to assist all college students, irrespective of path, obtain particular studying outcomes. Over 1,000 undergraduates enrolled in the curriculum.
Self-report survey outcomes point out that there have been no important variations in affective positive aspects by path. Students conveyed which elements of the curriculum have been crucial to their studying and improvement of research-oriented expertise. Students’ pursuits in biology elevated upon completion of the curriculum, inspiring a subset of CURE individuals to subsequently pursue additional analysis.
A rubric-guided efficiency analysis, employed to immediately measure studying, revealed variations in studying positive aspects for CURE versus ARE individuals, with proof suggesting a CURE can cut back the achievement hole between high-performing college students and their friends.
Catalyzing speedy discovery of gold-precipitating bacterial lineages with college college students.
Intriguing and probably commercially helpful microorganisms are discovered in our environment and new instruments enable us to find out about their genetic potential and evolutionary historical past. Engaging college students from completely different disciplines and programs in the seek for microbes requires an thrilling venture with progressive however simple procedures and targets.
Here we describe an interdisciplinary program to have interaction college students from completely different programs in the sampling, identification and evaluation of the DNA sequences of a distinctive but widespread microbe, Delftia spp.
A campus-wide problem was created to determine the prevalence of this genus, capable of precipitate gold, involving introductory degree environmental and life science programs, upper-level superior laboratory modules taken by undergraduate college students (juniors and seniors), graduate college students and workers from the campus.
The quantity of individuals concerned allowed for intensive sampling whereas undergraduate researchers and college students in lab-based programs participated in the pattern processing and analyses, serving to contextualize and solidify their studying of the molecular biology methods. The outcomes have been shared at every step by means of publicly accessible web sites and workshops.
This mannequin permits for the speedy discovery of Delftia presence and prevalence and is adaptable to completely different campuses and experimental questions.
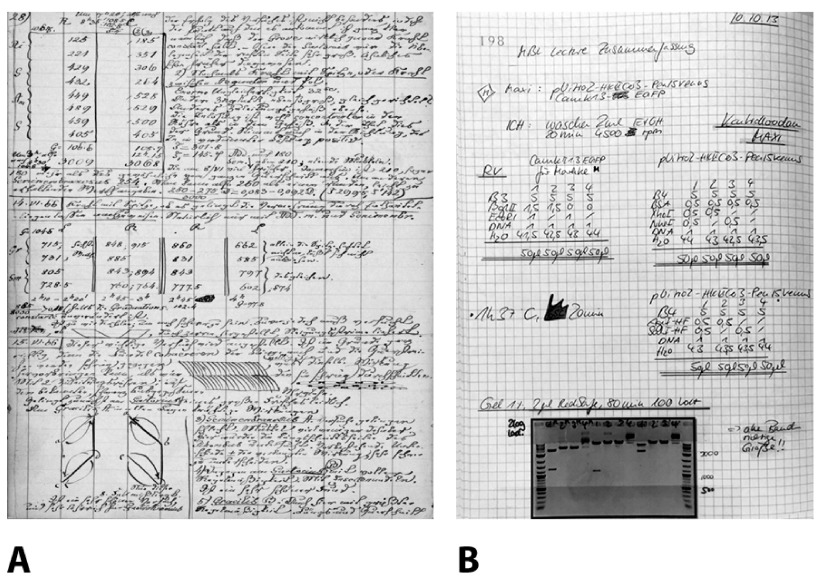
A pocket guide to electronic laboratory notebooks in the academic life sciences.
Every skilled doing lively analysis in the life sciences is required to hold a laboratory pocket book. However, whereas science has modified dramatically over the final centuries, laboratory notebooks have remained basically unchanged since pre-modern science.
We argue that the implementation of electronic laboratory notebooks (eLN) in academic analysis is overdue, and we offer researchers and their establishments with the background and sensible information to choose and provoke the implementation of an eLN in their laboratories.
In addition, we current knowledge from surveying biomedical researchers and technicians relating to which hypothetical options and functionalities they hope to see applied in an eLN, and which of them they regard as much less necessary.
We additionally current knowledge on acceptance and satisfaction of those that have not too long ago switched from paper laboratory pocket book to an eLN. We thus present solutions to the following questions: What does an electronic laboratory pocket book afford a biomedical researcher, what does it require, and the way ought to one go about implementing it?

(32)P measurment of urine samples and inside dose evaluation for radiation employees in life science laboratories.
(32)P measurements of urine samples and inside dose assessments had been performed for employees in life science laboratories. A process for pattern pre-treatment was established and validation was carried out to exclude interference and to detect (32)P ranges precisely.
The detection situations for Cherenkov radiation had been evaluated and the accuracy of Cherenkov radiation measurements validated. The analytical and measurement procedures had been utilized to urine samples collected from 11 employees from life sciences laboratories. The outcomes of the measurements typically indicated very low background radiation ranges, however every day urine samples from two employees had been above the minimal detectable exercise.
The (32)P concentrations for 2 of the employees had been 29.3 ± 10.4 Bq•d(-1) and 24.1 ± 11.8 Bq•d(-1), respectively, at consumption ranges of 4.12 okBq and a pair of.61 okBq. The efficient doses for these two employees had been 4.6 μSv and a pair of.9 μSv. Overall, the outcomes point out very low ranges of radioactivity, aside from instances associated to particular working situations.
Social science in a stem cell laboratory: what occurred when social and life sciences met.
We describe the expertise of conducting intensive social science analysis at the UK Stem Cell Bank from the viewpoint of each the particular person conducting the social science analysis and the Director of the Bank. We element the preliminary misunderstandings and issues held by each and the issues these precipitated.
Then we describe how the relationship developed as the challenge progressed and shared advantages turned obvious. Finally, whereas acknowledging potential areas of stress between the life and social sciences, we propose additional interplay between the disciplines would show helpful for each and speculate as to how this can be achieved.
In the dialogue we establish a set of studying factors from our expertise and definitions of social science terminology that will assist to inform future engagements between life and social scientists.
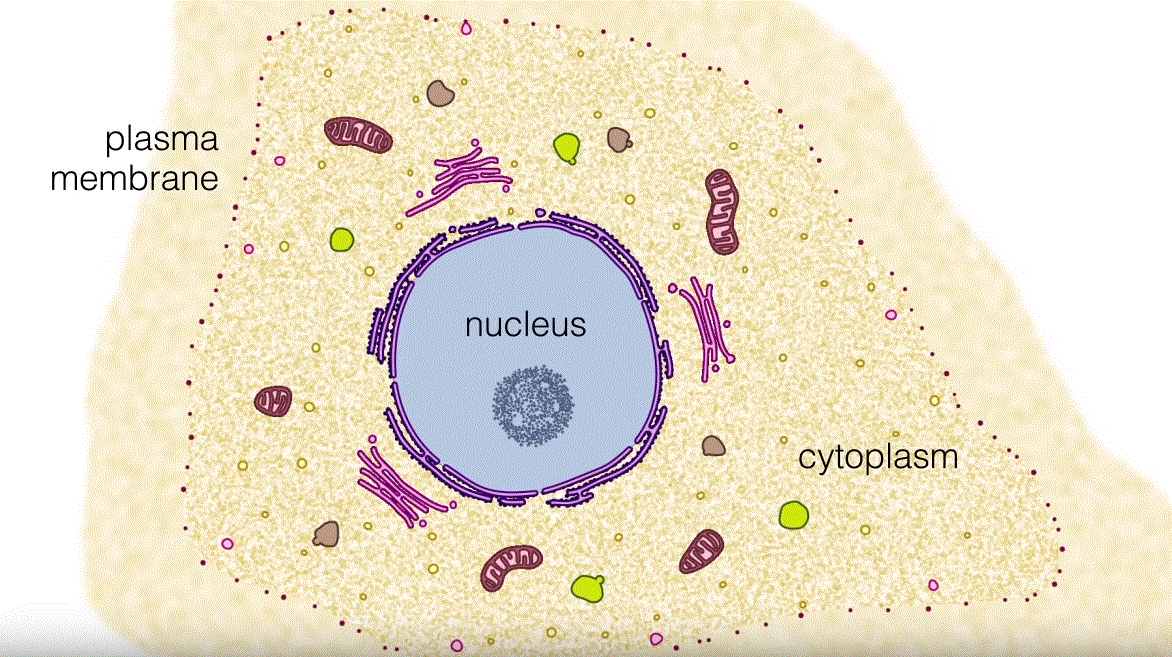
Mitochondria
Mitochondria are the nuclear power central of all living cells.
The role of Bcl2
The aforementioned experiments indicated that discerning BCL2 or BCL(X)L inhibition influenced on mitochondrial ATP production, but wasn’t sufficient to induce cell death.
In addition, AMPK, the significant mobile energy detector, was implicated in barrier function ( Peng et al., 2009; Wang et al., 2016) and also the boost in TEER may be a result of alterations in AMPK activity.
The smallest concentration of PA of 50 nM was inadequate to induce a reduction in TEER, while 100 nM of PA led to a delayed but comparable pattern as 200 nM of PA, which showed the biggest effects on Caco-2 monolayer permeability without detected cytotoxicity (Figure 3F). Along with 200 nM of PA also led to a considerable decline in the cellular energy status as reflected by decreased levels of ATP, currently at the stage where TEER was increased, but more conspicuous in the stage where TEER was diminished.
Mitochondria are critical cell organelles which do not just create the vast majority of mobile ATP but control cellular calcium homeostasis and modulate apoptotic pathways, one of a number of other important purposes:
- They’re also the principal source of intracellular reactive oxygen species (ROS)
- Throughout normal cell metabolism, ROS can operate as significant secondary messengers and there’s a balance between ROS production and their detoxification by cellular antioxidant systems.
- The dysfunctional mitochondria, marked by decreased ATP production and an increased production of ROS, disturb this equilibrium and have been theorized to contribute to aging and the evolution of age-related diseases.
In a vicious cycle, aberrant mitochondrial ROS cause additional damage to mitochondrial DNA (mtDNA), membrane lipids, and proteins, raising mitochondrial damage and additional bettering ROS leakage. Mitochondrial permeability transition pore (MPTP) opening leads to different pathologies between necrotic cell death after ischemic injuries or adrenal gland and brain ailments.
The MPTP happens over the inner mitochondrial membrane in which it’s permeable to solutes up to 1.5 kDa, and it opens into the existence of top matrix Ca2+ or reactive oxygen species.
Although regulators of these MPTP have already been identified, the molecular part of the MPTP remains.
But this version was challenged from the report which the mouse liver mitochondria lacking the enzymes Ant1 and Ant2 still failed mitochondrial permeability transition (MPT), albeit at considerably greater rates of Ca2+.
We investigated if the next Ant gene in mice, Ant4, compensates for the reduction of Ant1 and Ant2 and if mitochondria completely null for many ANT isoforms would experience MPT people comprise four Ant genes (Ant1 into Ant4), whereas mice deficiency Ant.
More recently, a version where the adrenal F1FO ATP synthase (ATPase) functions as the MPTP was suggested.
Cell Death
Stimuli may activate cell death, including the program, DNA damage, endoplasmic reticulum stress, growth factor and nutrient depletion disease and cognitive stress.
As part of their ever-growing household of atomic co-regulators, Pgc-1α can trigger a large group of genes and modulate the expression levels of enzymes involved in energy metabolism in response to signaling pathways which mediate thermogenesis, gluconeogenesis, muscle fiber type switching and mitochondrial biogenesis.
All these co-regulators exist and operate in big multi-protein complexes, where instead of simply binding to DNA, they govern Nrf-1/two and Tfam and regulate their transcriptional effectiveness by encouraging the following biochemical connections needed for induction or repression of gene transcription.
Additionally, Nrf-1/two also indirectly regulates the expression of adrenal DNA-encoded genes by potently causing the nuclear-encoded Tfam A, B1 and B2 (Tfam, Tfb1m and Tfb2m( respectively), that would be the regulators of their transcription and replication of the mitochondrial genome.
Past studies have shown that NaN is an inhibitor of the respiratory chain complex IV, which is affected in mitochondrial disorders.
In the current study, the term Pgc-1α indicate cascade, such as Pgc-1α household proteins (Pgc-1α, Nrf-1, Nrf-2 and Tfam) and Cox IV, was examined in PC12 cells to confirm if the signaling events were included NaN3-induced apoptosis.
Mitochondria are a significant source of ATP
In most cell types such as HeLa cells, as inhibiting mitochondrial ATP generation in these cells (using the ATP synthase inhibitor oligomycin) contributes to a 50 percent decline in total cellular ATP levels.
Third, treatment that was DMEM Acidic improved in cellular ATP levels.
Treating cells overall cellular ATP levels. The effect of absence of subunit α, Tb7760, or Tb2930 about the ATP synthase meeting was further analyzed by BN PAGE evaluation of dodecyl-maltoside-solubilized mitochondria in conjunction with activity-based discoloration, as explained above.
For its parental 29.13 cells and for uninduced controls, the ATP synthase was discovered as monomer, putative dimer, and free F1 contaminants, as observed in Figure 3 To its subunit α-silenced cells, no ATPase activity was discovered.
In such α and β RNAi cells in which decreased levels of the ATP synthase led to a reduction in membrane potential, the cells have been probably decreased in the capability to maintain functional requirements such as distinction, resulting in a reduction in growth rate. The ATP synthase, in all life cycle phases, may be discovered compared to the cytochromes but is most abundant during the stage, in which it works in ATP production through phosphorylation.
ATP synthase in bloodstream
Trypanosomes accounts for the creation of their membrane potential by hydrolyzing ATP made by substrate level phosphorylation.
The mechanism by which this reversible receptor switches in the ATP artificial into the ATP hydrolytic action remains not well known, but in E. coli, yeast, and mammalian cells it’s been indicated to be a reply to the proton motive force and the ADP/ATP equilibrium.
Next, ATP concentrations were quantified by us at the matrix utilizing the Lucm build.
In total moderate, wild-type cells revealed a top matrix ATP material (223 μM) which has been more than twice the amount detected in the cytosol ( Figure 4A). On the flip side, MELAS, NARP, and ρ0 cells had ATP degrees (101-112 μM) which were basically equal to the various cytosolic concentrations.
At OXPHOS-only” medium containing pyruvate as the only energy substrate (i.e.cells were made to use OXPHOS to create ATP), there was a decrease in total ATP degrees in all four cell lines.
But, there was a 45% decrease from the ATP content of wild-type cells as well as the value in full medium, whereas NARP cells, that include a partial defect in mitochondrial ATP synthesis, showed a marked (66 percent ) decrease in overall ATP.
Mitochondria is a significant double-membraned organelle that generates energy via bio-oxidation from eukaryotic cells.
Underneath SCI ailments, the first injury that led to mitochondria disruption causes loss of adenosine triphosphate (ATP) synthesis and raises several reactive oxygen species (ROS).
The capacity to revive mitochondria homeostasis and fix damaged mitochondria were outside the neutralizing capacities of antioxidant processes, inducing delayed neuronal apoptosis and necrosis.
The prior studies reported that SCI can cause adrenal morphological adjustments, cognitive impairment, and apoptosis, which might correlate with the pathological mechanisms of secondary damage after SCI 18, 19 Lately, studies report the use of mitochondria extends past energy generation to influence the two neural homeostasis and neurodegenerative processes in the central nervous system (CNS).
A potentially effective therapeutic strategy for SCI would be to revive mitochondria role in the spinal cord, enhancing neuron apoptosis and operational recovery. Proteins of the BCL2 family control the procedure of mitochondrial outer membrane permeabilization (MOMP) through apoptosis.
But, BCL2 proteins also modulate the bioenergetics status of cells throughout their management of mitochondrial fusion and fission dynamics, also may act directly to the mitochondrial respiratory chain.
For instance, BCL2 overexpression in human leukaemia cells enhanced oxygen consumption and mitochondrial respiration 15 In neurons, a pool of their anti-apoptotic BCL(X)L protein was found to localise from the inner mitochondrial membrane (IMM) and also to socialize with F1F0 ATP synthase, raising its enzymatic action and strengthening mitochondrial membrane potential (ΔΨm).
A truncated form of this anti-apoptotic MCL1 protein was shown to localize into the mitochondrial matrix and also to raise mitochondrial respiration.
Since anti-apoptotic BCL2 family proteins confer resistance of cancer cells to treatment, many selective inhibitors which target multiple or individual anti-apoptotic BCL2 proteins, have been developed.
The initial BCL2 selective inhibitor Venetoclax (ABT199) was accepted for clinical use for the treatment of chronic lymphocytic leukemia using 17p deletion.
But little information is now accessible if selective inhibition of BCL2 or alternative anti-apoptotic BCL-2 protein for example BCL(X)L additionally affects cognitive functioning and energetics in cancer cells.
Logevity
The experiments using metformin and FCCP, implicate the participation of a practical ETC into MondoA transcriptional activity.
In statistics we elected to not present, we utilized 143Brho cells paired with mitochondria using mutations in cytochrome B (complex 3) to show that the ETC need for lies of complicated 3 and so upstream of ATP synthase. Even though a short-term introduction of the mPTP seems to serve as a standard calcium-release mechanism that’s needed for appropriate metabolic regulation, irreversible formation and consequent opening of the mPTP are crucial aspects in adrenal dysfunction and mitochondria-driven cell death.
When the mitochondria are exposed to elevated concentrations of calcium, then they also experience an immense and permanent swelling which contributes to a sudden increase in permeability to small solutes of the IMM, abolishing the chemiosmotic gradient throughout the IMM 29, which then uncouples OXPHOS, resulting in a drop in ATP production and a rise in ROS formation.
Further rupture of the outer mitochondrial membrane leads to the extrusion of both cytochrome c, an integral step in the initiation of apoptosis.
The mPTP can also play a part in the regulation of energy generation because of the double purpose of the ATP synthase in the ATP manufacturing and mPTP formation.
ROS generation may initiate diverse cellular responses, including tripping signaling pathways involved in cell security, initiating coordinated activation of mitochondrial fission and autophagy to maximize removal of abnormal mitochondria and cells, also ensuring that the harm doesn’t propagate to neighboring mitochondria and cells.
The two elevated levels of ROS (oxidative stress) and too low levels of ROS (reductive stress) are dangerous and might play causative roles in the pathologies about the remarkable reversal of redox surroundings.
Ros, reactive oxygen
Excessive ROS generation from the center under pathophysiological conditions contributes to mitochondrial dysfunction and bioenergetic reduction and leads to numerous mobile pathologies from the center. We observed an gain in the degree of those autophagy-related proteins, Sequestosome1/p62 (p62) along with also the lipidated form of this microtubule-associated protein 1 light chain 3 (LC3), also LC3-II, 24 hours following the accession of AA, implying the regulation of autophagy.
This confirms the hypothesis which AA-treated cells utilized to clean restrict toxicity and the mitochondria. Unsurprisingly, Aa treatment caused a radical and speedy decrease in mitochondrial respiration, verifying that the reduction of adrenal function .
With no energy generation by OXPHOS and so as to facilitate mobile survival, ARPE-19 cells had triggered glycolysis pathways, as evidenced by a rise in ECAR.